Principal Investigator:
Dan Avi Landau, Bibliowicz Family Professor of Medicine
Background & Unmet Need
- Circulating tumor DNA (ctDNA) in plasma cell-free DNA (cfDNA) enables non-invasive cancer detection through routine blood draws
- ctDNA levels correlate with tumor burden and reflect responses to treatment, serving as a valuable biomarker for early detection and disease monitoring
- While late-stage cancers show tumor fractions (TFs) up to 20% in cfDNA, early-stage disease exhibits dramatically lower TFs
- A conventional blood draw (5mL) yields only 2,000-5,000 genomic equivalents of cfDNA, limiting sequencing depth
- The combination of low TF and limited genomic equivalents creates a fundamental barrier to early cancer detection through conventional mutation sequencing
- Unmet Need: Development of ultra-sensitive methods capable of detecting plasma tumor fraction <0.1% to enable early cancer detection
Technology Overview
- The Technology: MRDetect, a tumor-informed method for ultra-sensitive ctDNA detection using genome-wide mutational integration and an advanced error suppression framework
- MRDetect is available via non-exclusive license
- MRDetect uses average depth whole genome sequencing (WGS) to pool information across multiple sites in the genome, increasing the effective ceiling of sequencing depth
- Read-level error suppression frameworks address noise associated with lower quality sequencing metrics and read depth in single nucleotide variants (SNVs) and copy number variants (CNVs)
- PoC Data: MRDetect enables ctDNA detection in fractions as low as 10-5 with a modest sequencing depth (35x), with assay-level specificity of 95%
- PoC data has been generated in clinical plasma samples from patients with lung adenocarcinoma, colorectal cancer, and metastatic melanoma
Technology Applications
- Early detection of cancer in high-risk populations
- Detection of minimal residual disease following cancer treatment
- Patient stratification for adjuvants to reduce unnecessary treatments
- Treatment selection based on patient-specific molecular features and response monitoring
Technology Advantages
- Simple WGS workflow does not require custom panel creation or molecular barcodes and can work with limited input material
- Enables dynamic treatment optimization through ongoing monitoring of patient response and outcomes
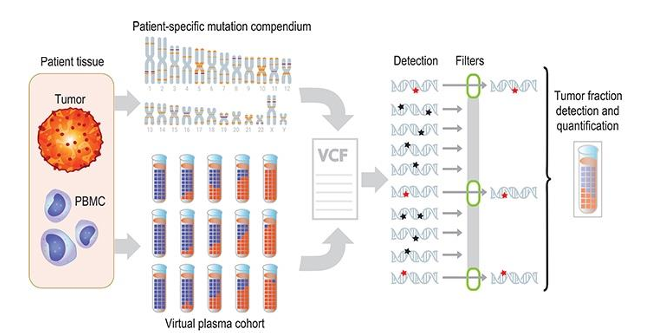
Figure: Illustration of genome-wide SNV integration for inference of plasma TF, and analytical validation scheme.
Resources
Intellectual Property
Patents
- US20210043275A1: "Ultra-sensitive detection of circulating tumor dna through genome-wide integration"
- EP3759237A4:"Ultra-sensitive detection of circulating tumor dna through genome-wide integration"
- US20210002728A1: "Systems and methods for detection of residual disease"
- EP3759238A4: "Systems and methods for detection of residual diseases"
- Issued patents in SG, AU, CN, JP
- Additional Applications in CA, JP, IN, IL, KR, HK, SG, CN, AU, US
Cornell Reference
- 7874
Contact Information

For additional information please contact
Jamie Brisbois
Manager, Business Development and Licensing
Phone: (646) 921-4743
Email: jamie.brisbois@cornell.edu

